Identification of poorly resolved sequencing traces
- The trace peaks are very broad or “blurry”. This blurriness occurs before base 500 for trace collected using 36cm capillary arrays, or before base 650 for traces collected on 50cm capillary arrays.
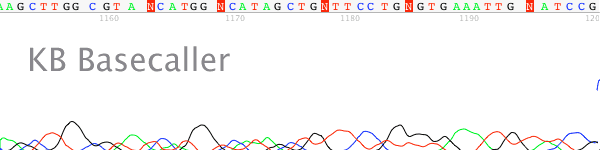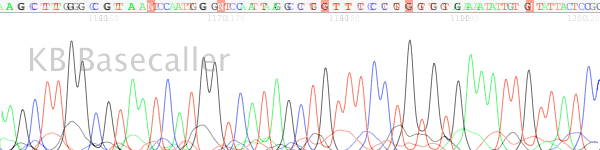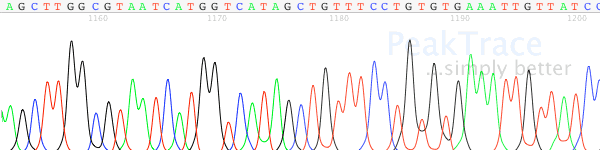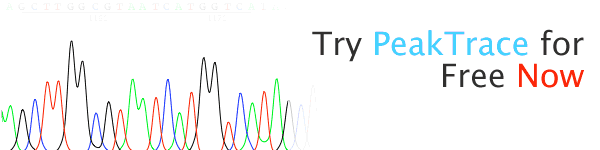
- The nucleotide runs in the trace sequence appear as indistinct blobs rather than as individually separate peaks (Figure 1).
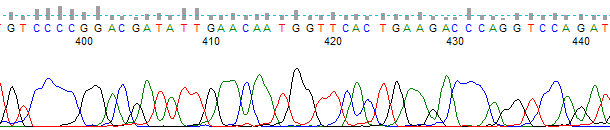
Figure 1. Example of a “blurry” or poorly resolved trace chromatogram collected using a ABI 3100 50cm array.
Causes of blurry sequencing traces
- Capillary overload. This is normally caused by running “dirty” samples with large amounts of template (or other) DNA, proteins or salt.
- High sequencing run voltages. The higher the run voltage the worse the resolution of the peaks in the trace file. High run voltages can also cause problems for the KB base caller resulting from the collection of the void peak.
- Old array. The capillary array only has a certain lifetime (ABI only guarantees the arrays for 300 runs). After around 1000 runs, the inside of the capillaries in the array become coated in “junk” (proteins mostly) that are not washed out when the polymer is replaced. This junk coating interacts with the DNA fragments during a run reducing the peak resolution.
Solving blurry sequencing trace problems
- Use clean DNA. Avoid using templates that contain a significant amount of Genomic DNA, protein or salt.
- Check the concentration of the template.If the template concentration is too high then re-sequence using less template.
- Check that the correct run voltage was used. This can be determined by examining the run protocol contained with the ABI trace file.
- Change the array. If the array has been used of over 1000 runs then it may need to be replaced.
- Use the PeakTrace Basecaller. While using the PeakTrace Basecaller does not solve the underlying production problem that leads to poorly resolved traces, it is able to provide a much better resolved trace than the KB Basecaller. It is always best to fixed the production problem that causes poorly resolved traces, but PeakTrace can help rescue traces that have already been collected.
For more information on detecting DNA sequencing trace problems please visit the QualTrace III DNA sequencing analysis software page.
Return to the main DNA sequencing troubleshooting page.