Developer: Tom Hall
Supported platforms: Windows 95 and higher
Download: http://www.mbio.ncsu.edu/BioEdit/bioedit.html
Features
View Quality scores
No
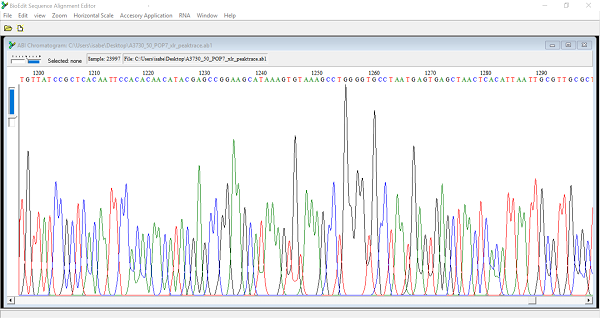
View raw data
Yes
Assembly
Yes, but you have to manually align up each sequence and manually create a consensus sequence (which other programs are very good at).
Input file formats
ABI files (.abi and .ab1), FASTA, Genbank, Fasta, Phylip 3.2, Phylip 4, GCG, Clustal, and NBRF/PIR formats (a complete list can be found in the manual).
Output file formats
ABI files, text files, FASTA, Genbank, Fasta, Phylip 3.2, Phylip 4, GCG, Clustal, and NBRF/PIR formats (a complete list can be found in the manual).
Trimming
No trimming of low quality regions
Integrated Blast
Yes
Open multiple windows
Yes
Edit bases
Yes
ABI limits (regions outside of clear range region are displayed in grey)
No
Other functions
- Reverse complement sequence
- Rudimentary phylogenetic tree viewer
- Plasmid drawing and restriction site mapping
- Translation
- ClustalW multiple sequence alignment
A complete list can be found on the BioEdit website
Cost
Free, but no longer maintained. The website seems to also be down these days. We have a copy of the installer if anyone wants to get a copy.
Comments
BioEdit is free and provides many basic functions for protein and nucleic sequence editing, alignment, manipulation and analysis. The software works is no longer maintained. If you only need a trace viewer or want to do basic editing other free software options are better because they have more trimming options and display the quality values (phred scores). However, if you are looking for a free alignment program BioEdit might be right for you.