Developer: Nucleobytes
Supported platforms: MacOSX 10.7 or later
Download: https://nucleobytes.com/4peaks/index.html
Features
View Quality scores
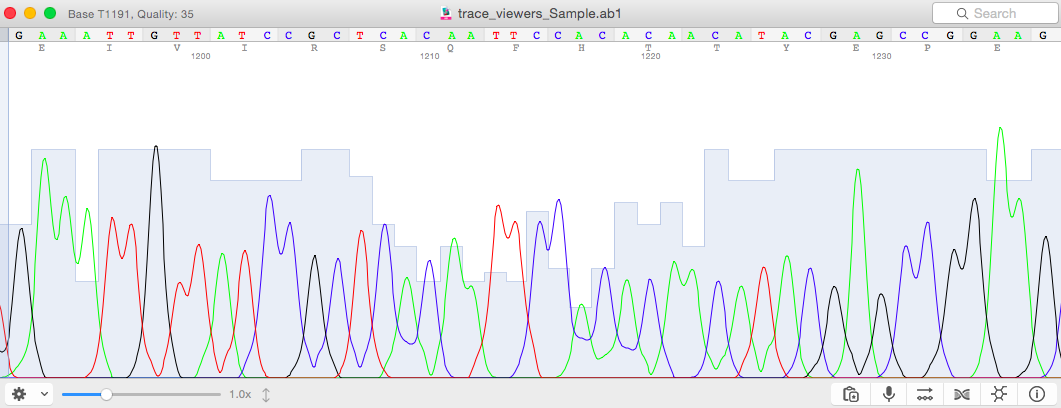
Yes. Quality scores are presented as blue bars behind the peaks and at the top of the window when a peak is selected.
View raw data
No. There is now way to view the raw data in 4Peaks.
Assembly
No. 4Peaks does not contain any features for DNA alignment or assembly.
Input file formats
ABI, SCF, ALF, PLN, EXP, ZTR, BIO and TXT.
Output file formats
SCF, PLN, ZTR.
Trimming
Yes. Through the quality overview function you can trim below a set quality value. It is possible to manually select areas to trim at the 3′ and 5′ ends.
Integrated Blast
Yes. You can run a BLAST search through 4Peaks. However, it will take you into a separate web browser.
Open multiple windows
Yes.
Edit bases
It is possible to edit, insert and delete bases in 4Peaks.
ABI limits (regions outside of clear range region are displayed in grey)
No. This function is not available in 4Peaks.
Other functions
The program is full of nifty features, you have the ability to open a separate txt window containing the sequence and select from the peaks/txt file at the same time to compare specific regions. You also have the ability to translate the DNA sequence into the amino acid sequence across 3 separate frames. One handy feature is the speech function where 4Peaks will verbally read out the sequence of the file so you can follow along while looking at another sequence to compare differences.
Cost
4Peaks is free.
Comments
4Peaks boasts that it is able to “render peaks better than anyone else”. Traces appear sharper when viewed through 4Peaks compared to other chromatogram viewing programs, which allows for a easy viewing. A major downside of 4Peaks is that the quality score bars are shown in the background which creates confusion when viewing the quality and chromatogram in the same window. There is also no way to view the raw data signal.