Developer: GSL Biotech LLC
Supported platforms: Windows 7 or later, MacOS 10.8 or later, Ubuntu Linux 14.04 or later, and Fedora Linux 21 or later
Download: https://www.snapgene.com/snapgene-viewer/
Features
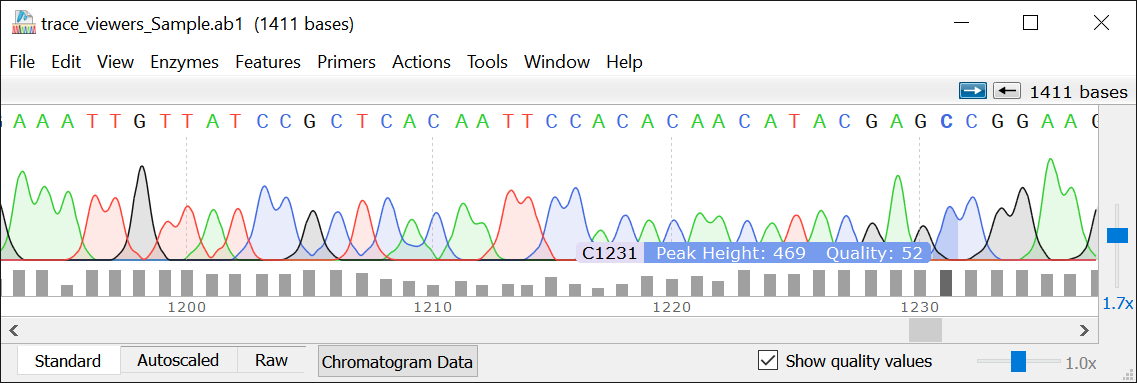
View Quality scores
Quality scores can be toggled on or off. They appear at the bottom of the trace, and at the location of the cursor when hovering over a peak.
View raw data
Raw data can be viewed through a tab at the bottom of the window, you can also find a tab for information on the chromatogram here as well.
Assembly
No. Only If you upgrade to the paid version ‘Snap Gene’.
Input file formats
Snap Gene Viewer can handle .ab1, .ztr and SCF chromatograms. It can also input sequence files from over 10 different programs including Geneious, GenBank and DNA Dynamo.
Output file formats:
SCF, ZTR and FASTQ.
Integrated Blast
There is an option to run a BLAST alignment through the viewer but it will take you out of the program to an internet browsing window.
Open multiple windows
Yes.
Edit bases
Yes.
ABI limits (regions outside of clear range region are displayed in gray)
No.
Other Functions
Snap Gene Viewer is the base level for the paid Snap Gene molecular cloning software. Sequence features can be automatically annotated within the program to identify features such as primer locations, restriction sites, ORF’s and ligation sites. Cloning processes have also been simulated with support for techniques such as Gibson assembly and Gateway cloning.
Cost
Snap Gene Viewer is free however they have multiple plans if you would like to upgrade to the premium Snap Gene program, starting at $345 for academics or $1,245 for commercial use.
Comments
Snap Gene Viewer is simple to use and displays the data in a somewhat unusual and colorful manner where the entire peak is colored rather than just the outline. The simplicity of the program is a real winning point with this viewer.