UniSeq primer walking DNA sequencing system manual
UniSeq is a universal, innovative and robust system for the sequencing of DNA. UniSeq is completely compatible with current DNA sequencing technology based on the Sanger method and can be incorporated into existing protocols.
The UniSeq system uses two specifically designed primers in each sequencing reaction selected from a presynthesised, 768-member oligonucleotide library of E- and T-primers. The UniSelect software is used to select the optimal E- and T-primer pairs that are added to the sequencing reaction. The DNA polymerase present in the sequencing reaction catalyses the production of a single template specific primer (S-primer). By the combinatorial application of the E- and T-primers from the UniSeq library over 131,000 different and unique S-primers can be generated – sufficient to allow sequencing of nearly all DNA sequences. The overall process is illustrated in Figure 1.
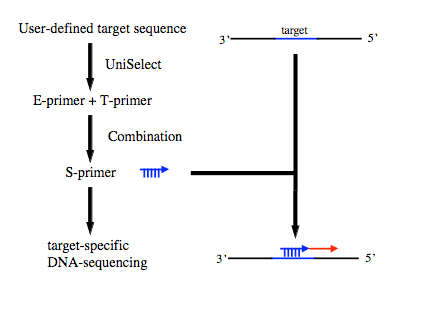
Figure 1. The UniSeq process
UniSeq prime kit components
The UniSeq primer library
The UniSeq primer library consists of 768 individual primers. These primers are arrayed into two 384-well microtitre plates and are shipped dry. Dry primers are stable at 4 deg.C for at least 12 months. After reconstitution the UniSeq primer library should be stored at –20 deg.C.
The UniSelect software
The UniSelect software automates the process of selection the optimal E- and T-primers pairs for a user defined target sequence. The UniSelect software accepts PHRED generated PHD files and generates an output list with the target sequence index and the corresponding two primers indices. For further details on the software please see the UniSelect User Manual.
Sequencing reaction modifiers
UniSeq Reaction Modifier solutions are provided for most common sequencing chemistries (Table 1). Each reaction modifier is optimized for a particular sequencing chemistries. Nucleics does not recommend the use of sequencing modifiers for other chemistries.
The UniSeq Reaction Modifiers are provided as 20x stock solutions and should be long-termed stored at –20 deg.C. Excessive freezing and thawing of the reaction modifier should be avoided. The UniSeq Reaction modifier can be stored for up to 2 days at room temperature or 37 deg.C without any loss of activity.
Sequencing reaction enhancer
A UniSeq Reaction Enhancer is provided (Yellow-capped tube). The reaction enhancer is designed to improve the sequencing of difficult DNA templates such as plasmids with strong secondary structures or high GC-content. Nucleics does not recommend the use of other DNA sequencing additives such as betaine or dimethylsulfoxide (DMSO), as they can interfere with the UniSeq system. The use of the reaction enhancer is optional, however, certain difficult templates will sequence better in the presence of the enhancer.
The 8 x UniSeq Reaction Enhancer should be stored at 4 deg.C.
Table 1. UniSeq reaction modifiers supplied
UniSEQ Protocols
This section will provide protocols and information for the successful set up and use of the UniSeq system.
Preparation of the library
Before use, the UniSeq primer library must be resuspended in molecular biology grade water. Add the amount of nuclease-free molecular biology grade water indicated on the tray to each of the microtitre plate well. To allow the primers to redissolve incubate for 15 to 30 min at room temperature before use.The primer library should be stored at –20 deg.C when not being used.
Primer selection
Please see UniSelect manual.
UniSeq reaction
Nucleics recommend the use of half-volume (10µl) DNA sequencing reaction. If desired, full-scale reactions can be used with the system by doubling the volumes recommended in this protocol.
- Pipette 3µl of the each of the two UniSeq primers suggested by the UniSelect software into PCR tube or microtitre plate well.
- Dry the primers at 80 deg.C for ~20 min.
- Add to each tube/well with the dried primers the following reagents:
- 4µl of sequencing chemistry premix
- 0.5 µl of 20x UniSeq reaction modifier
- 1.25 µl of 2.5x UniSeq reaction enhancer (optional)
- up to 4.25 µl to 5.5µl of DNA template
- Molecular biology grade water to 10µl
4. Thermocycler tubes/microtitre plates using the following conditions:
- 40 cycles: 70 deg.C for 5 s; 32 deg.C for 5 s
- 1 cycle: 96 deg.C for 1 min
- 50 cycles: 96 deg.C for 10 s; 46 deg.C for 30 s (decrease by 0.1 deg.C per cycle); 60 deg.C for 4 min
5. Samples may be cleaned as per standard dye terminator sequencing reaction.
Recommendations
This section provides recommendations for the successful use of UniSeq.
Template concentration and quality
The success rate of UniSeq reactions is highly dependant on the DNA template concentration and quality. Nucleics recommends the use of 50 to 100 ng of DNA template per kilobase of template in each UniSeq reaction.
The DNA template should be of the highest quality and purity. For plasmid templates, Nucleics recommends the use of commercially available plasmid miniprep kits. If PCR products are used as templates excess primers, nucleotides and DNA polymerase should be removed.
Sequencing chemistry dilution
Nucleics is aware that many DNA sequencing user dilute the sequencing chemistry in order to save costs. Sequencing chemistry dilution is possible using UniSeq, however the user also has to be made aware that a reduced signal intensity and therefore reduced sequence quality might be obtained.
Nucleics recommends a maximal dilution of 1:1 for the BigDye terminator system with 2.5x Sequencing Dilution buffer (ie. in a 10µl sequencing reaction, 2µl of BigDye premix plus 2µl of 2.5x Dilution Buffer [200 mM Tris-HCl pH 9.0, 5 mM MgCl2] may be used). Other dilution buffers have not been tested and Nucleics does not guarantee the compatibility of these buffers with UniSeq.
Thermocycler and cycling conditions
UniSeq has been optimized for PCR thermocycler with temperature ramping ability (eg. the PerkinElmer GeneAmp 9600 and 9700). Other thermocycler without temperature ramping such as the Stratagene Gradient 96 Robocycler have not been extensively tested and may require modifications of the cycling protocol for optimal results.
Reaction clean-up
UniSeq reactions can be purified in the same fashion as normal dye terminator sequencing reactions and Nucleics recommends users follow the manufacturers recommendations for the sequencing chemistry and DNA sequencer used. Please note that UniSeq is incompatible with biotin/strepavidin-based clean-up systems.
Technical Support
Nucleics provides expert technical support for the UniSeq DNA sequencing software. Please email any questions or comments about UniSeq to our support team at {This email is obscured. Your must have javascript enabled to see it}.