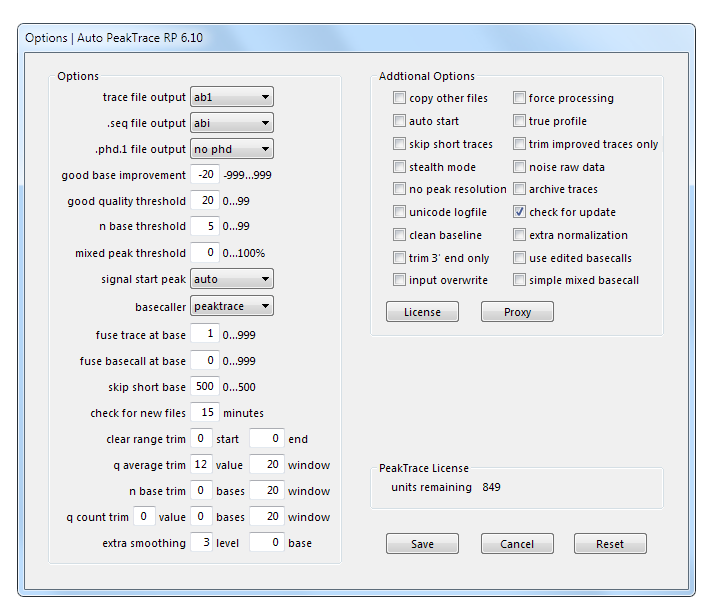
A question we get asked occasionally is what PeakTrace Basecaller settings are used with the free PeakTrace Online version? Because the online version is intended to support the widest range of sequencer run conditions, while showing (some) of the power of PeakTrace, it uses a set of conditions rather different to default values. The free version of PeakTrace uses the linux command line software with the following the options settings:
-ft 300 -c -40 -wq 12 20 -xb 7 -cb yes -bc pk -ts no -xs 1000 2 -sm 700
These values correspond to the Auto PeakTrace and Auto PeakTrace RP Option settings:
- fuse trace at base: 300
- good base improvement: −40 bases
- q average trim: 12 threshold; 20 base window
- clean baseline: checked
- extra baseline: 7
- basecaller: peaktrace/kb (PeakTrace Basecalling with KB Fallback)
- trim 3’ end only: checked
- extra smoothing: level 2; start base 1000
- skip short/pcr base: 700
These settings are not suitable if you are trying to detect polymorphic sites (i.e. mixed peaks). For this you need to use a paid version of PeakTrace and a mixed peak threshold above the default 0.
The PeakTrace default values are usually the best place to start, but you should always trial the various options to find what works best for your needs. If you would like further advice on the best settings to use with your sequencing data, please don’t hesitate to contact us @ {This email is obscured. Your must have javascript enabled to see it}.